We have developed a number of algorithms and pieces of software for analysing mass spectrometry data. Links can be found below, together with more details in the associated publications. More will be made available here soon, so please check back!
DynamXL
DynamXL accounts for the flexible nature of both protein and cross-linker molecules by (1) predicting the accessible space of amino acid side chains, (2) aggregating measures from multiple protein conformations and (3) measuring distances with a powerful shortest path algorithm, Theta*. The software, developed in Python, can be controlled via a graphical user interface, via terminal, or directly imported into an external code.
DynamXL has been developed by Matteo while in the Benesch group, at the University of Oxford. Further information and support can be found at Matteo Degiacomi's
website. EM∩IM
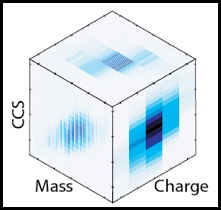
EM
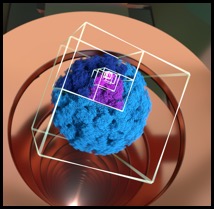
∩IM is software relating electron density maps, typically obtained from transmission electron microscopy (EM), with ion-mobility mass spectrometry (IM-MS) data.
It allows the user to estimate collision cross sections from density maps, and thereby enabling the mutual correlation and validation of IM-MS and TEM. Further, the software provides a means to define an appropriate EM threshold value ("contour"), leading to a density map visualisation more representative of actual data. EM
∩IM can be controlled by a Graphical User interface or interactively via Python commands.
DownloadFurther information and support can be found at Matteo Degiacomi's
website. IMPACT
Ion Mobility Projection Approximation Calculation Tool (IMPACT) is a computational tool for calculating collision cross-sections from structural models. It is designed for the high-throughput processing of large molecular structures and models (>10,000 atoms) without significant decrease in accuracy. It can accept input coordinate files for single structures (e.g. from X-ray crystallography), ensembles (e.g. from NMR), electron density maps (e.g. from electron microscopy), and coarse-grained models (e.g. from SAXS). IMPACT is 2 to 8 orders of magnitude faster than other algorithms, which have typically been designed for much smaller molecules, and can be invoked directly as a command-line tool in UNIX/Linux or Mac OSX, or used as a library linked with other software for molecular dynamics and integrated structural biology applications.
DownloadFurther support and for enquiries, please contact
Erik Marklund at Uppsalla University. UniDec
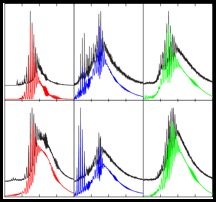
Universal Deconvolution (UniDec) is software to deconvolute both simple and complex nanoelectrospray mass spectra of proteins and the assemblies they form. UniDec was developed largely in the
Robinson Group, building on initial work by Mike Marty when he was at Washington University, and incorporates elements from CHAMP and CHAMPIon, and has a user-friendly graphical interface. In manuscript we describe how UniDec can enable the quantitative interpretation of mass spectra from small proteins to heterogeneous nanodisc assemblies, while allowing the extraction of biophysical quantities including rate constants and binding affinities.
DownloadFor support and enquiries, please contact
Mike Marty at University of Arizona.
CHAMP
Calculating the Heterogeneous Assembly and Mass spectra of Proteins (CHAMP) is software to deconvolute complex nanoelectrospray mass spectra arising from mixtures of intact protein assemblies. Our approach is based on an empirical understanding of what governs the appearance and variations in the mass spectra of protein assemblies. This is complemented by an automated and methodical data fitting algorithm to objectively identify the constituents of a polydisperse protein ensemble and to quantify their relative abundance. In our initial study
we employed CHAMP to deconvolute the complexes formed between a plant sHSP and different target proteins. CHAMP is available on request, but most users will find
UniDec more appropriate for their needs.
CHAMPIon
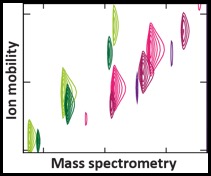
CHAMPIon is an extension of
CHAMP to include the separation in size (as well as mass) afforded by ion mobility mass spectrometry experiments. The additional dimension of separation is very useful in reducing the ambiguity of assigning peaks in mass spectra, and has allowed us to accurately determine
dissociation constants of ligand binding to protein assemblies. In addition, through exploiting the arrival-time information obtained, it is possible to
distinguish between models to determine likely structures of the protein assemblies in question. CHAMPIon is available on request, but most users will find
UniDec more appropriate for their needs.
Polymake
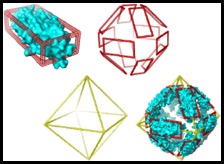
This algorithm allows the user to build candidate oligomer structures based on a protomeric building block and polyhedral or ring-like scaffolds. Spatial restraints to guide the assembly of the oligomers can be included, and the algorithm also incorporates a modified version of
Mobcal, enabling the rapid estimation of collision cross section for direct comparison to ion mobility data.
We used Polymake to construct models for the sHSP
αB-crystallin, and by filtering them relative to our experimental data, revealed the likely architecture of this polydisperse protein .
DrOP
Droplet Occupancy Predictor (DrOP) is an algorithm designed together with Giorgio Favrin to predict the expected extent of nonspecific protein aggregation during the electrospray ionisation mass spectrometry experiment. This monte-carlo algorithm combines the (measured) concentration of a given analyte in solution, together with empirical knowledge of the droplet sizes in electrospray ionisation.
DrOP is useful when examining systems with equilibrium dissociation constants at micromolar or above.
Excel Utilities
Over the years we have written a number of spreadsheets to provide some basic aids in assigning, assessing, and interpreting ion mobility and mass spectrometry data. While these have been largely rendered obsolete with the development of more sophisticated software solutions, they can still be useful in certain cases, and are available for download here. These are: Mass and Charge State Evaluation and Determination (
MaCSED), which aids in determining the correct charge state assignment for an electrospray spectrum; Gaussian Fitting of Arrival Times (
GFAT), which enables improved extraction of ion mobility peak centroids and widths; and Expected Ion Mobility Distribution (
EIMD) which helps you estimate whether conformations are separable in an ion mobility experiment.